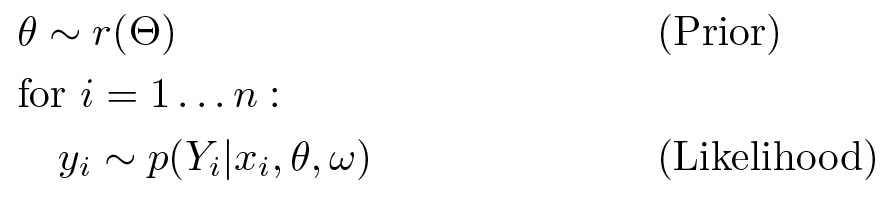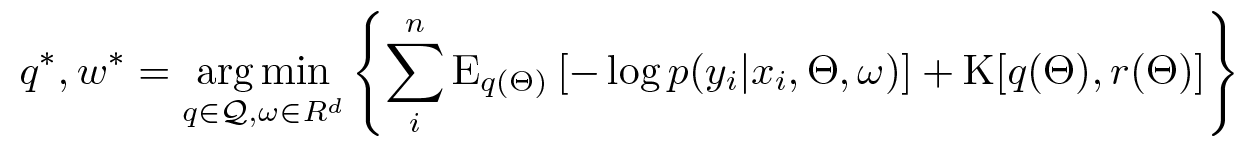Posted by Emily Fertig, Joshua V. Dillon, Wynn Vonnegut, Dave Moore, and the TensorFlow Probability team
In this post, we introduce new tools for variational inference with joint distributions in TensorFlow Probability, and show how to use them to estimate Bayesian credible intervals for weights in a regression model.
Overview
Variational Inference (VI) casts approximate Bayesian inference as an optimization problem and seeks a ‘surrogate’ posterior distribution that minimizes the KL divergence with the true posterior. Gradient-based VI is often faster than MCMC methods, composes naturally with optimization of model parameters, and provides a lower bound on model evidence that can be used directly for model comparison, convergence diagnosis, and composable inference.
TensorFlow Probability (TFP) offers tools for fast, flexible, and scalable VI that fit naturally into the TFP stack. These tools enable the construction of surrogate posteriors with covariance structures induced by linear transformations or normalizing flows.
VI can be used to estimate Bayesian credible intervals for parameters of a regression model to estimate the effects of various treatments or observed features on an outcome of interest. Credible intervals bound the values of an unobserved parameter with a certain probability, according to the posterior distribution of the parameter conditioned on observed data and given an assumption on the parameter’s prior distribution.
In this post, we demonstrate how to use VI to obtain credible intervals for parameters of a Bayesian linear regression model for radon levels measured in homes (using Gelman et al.’s (2007) Radon dataset; see similar examples in Stan). We demonstrate how TFP JointDistributions combine with bijectors to build and fit two types of expressive surrogate posteriors:
- a standard Normal distribution transformed by a block matrix. The matrix may reflect independence among some components of the posterior and dependence among others, relaxing the assumption of a mean-field or full-covariance posterior.
- a more complex, higher-capacity inverse autoregressive flow.
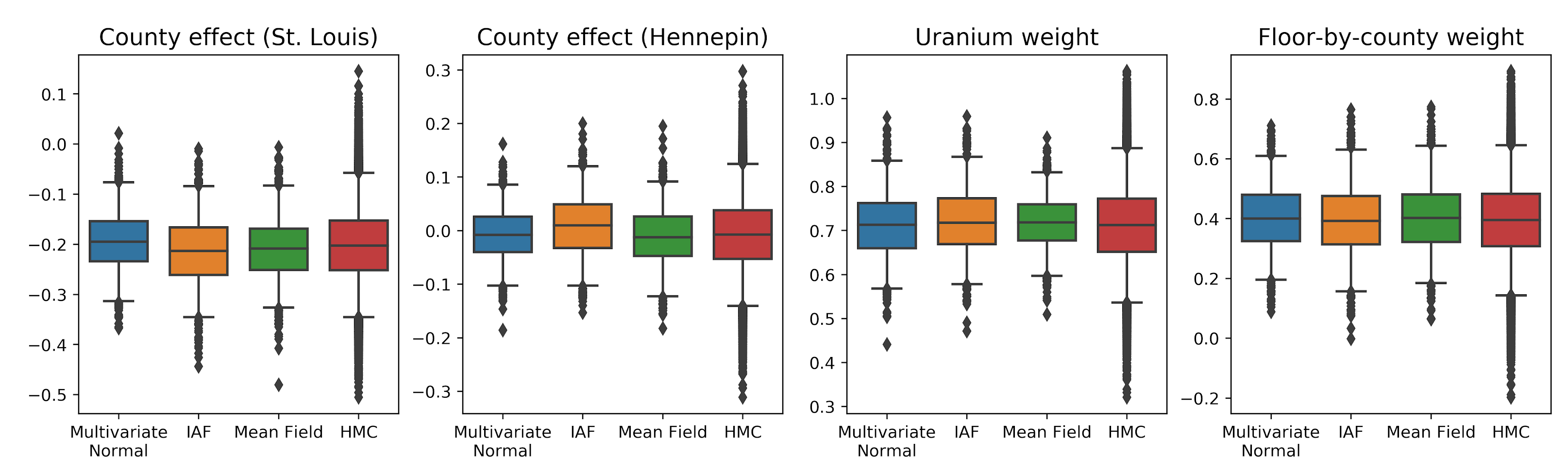
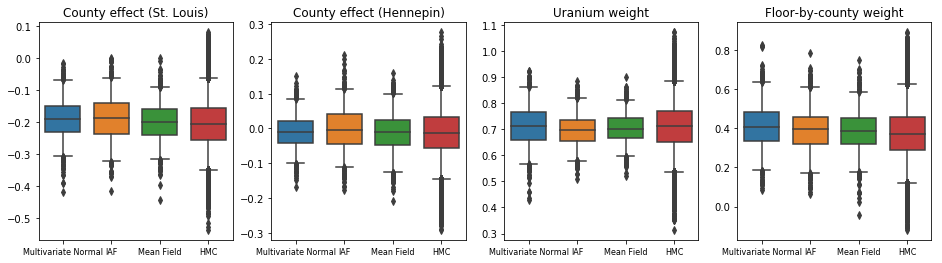
The surrogate posteriors are trained and compared with results from a mean-field surrogate posterior baseline. These plots show credible intervals for four model parameters obtained with the three VI surrogate posteriors, as you’ll learn about below, as well as Hamiltonian Monte Carlo (HMC) for comparison.
You can follow along and see all the details in this Google Colab.
Example: Bayesian hierarchical linear regression on Radon measurements
Radon is a radioactive gas that enters homes through contact points with the ground. It is a carcinogen that is the primary cause of lung cancer in non-smokers. Radon levels vary greatly from household to household.
The EPA did a study of radon levels in 80,000 houses. Two important predictors are:
- Floor on which the measurement was taken (radon higher in basements)
- County uranium level (positive correlation with radon levels)
Predicting radon levels in houses grouped by county is a classic problem in Bayesian hierarchical modeling, introduced by Gelman and Hill (2006). We are interested in credible intervals for the effect of location (county) on the radon level of houses in Minnesota. In order to isolate this effect, the effects of floor and uranium level are also included in the model. Additionally, we will incorporate a contextual effect corresponding to the mean floor on which the measurement was taken, by county, so that if there is variation among counties of the floor on which the measurements were taken, this is not attributed to the county effect.
The regression model is specified as follows:
in which i indexes the observations and countyi is the county in which the ith observation was taken.
We use a county-level random effect to capture geographical variation. The parameters uranium_weight and county_floor_weight are modeled probabilistically, and floor_weight and the constant bias are deterministic. These modeling choices are largely arbitrary, and are made for the purpose of demonstrating VI on a probabilistic model of reasonable complexity. For a more thorough discussion of multilevel modeling with fixed and random effects in TFP, using the radon dataset, see Multilevel Modeling Primer and Fitting Generalized Linear Mixed-effects Models Using Variational Inference.
The full code for this example is available on Github.
Variables are defined for the deterministic parameters and the Normal distribution scale parameters, with the latter constrained to be positive.
import tensorflow as tf
import tensorflow_probability as tfp
tfd = tfp.distributions
tfb = tfp.bijectors
floor_weight = tf.Variable(0.)
bias = tf.Variable(0.)
log_radon_scale = tfp.util.TransformedVariable(1., tfb.Exp())
county_effect_scale = tfp.util.TransformedVariable(1., tfb.Exp())We specify the probabilistic graphical model for the regression as a TFP JointDistribution.
@tfd.JointDistributionCoroutineAutoBatched
def model():
uranium_weight = yield tfd.Normal(0., scale=1., name='uranium_weight')
county_floor_weight = yield tfd.Normal(
0., scale=1., name='county_floor_weight')
county_effect = yield tfd.Sample(
tfd.Normal(0., scale=county_effect_scale),
sample_shape=[num_counties], name='county_effect')
yield tfd.Normal(
loc=(log_uranium * uranium_weight
+ floor_of_house * floor_weight
+ floor_by_county * county_floor_weight
+ tf.gather(county_effect, county, axis=-1)
+ bias),
scale=log_radon_scale[..., tf.newaxis],
name='log_radon')We pin log_radon to the observed radon data to model the unnormalized posterior.
target_model = model.experimental_pin(log_radon=log_radon)Quick summary of Bayesian Variational Inference
Suppose we have the following generative process, where 𝜃 represents random parameters (uranium_weight, county_floor_weight, and county_effect in the regression model) and ω represents deterministic parameters (floor_weight, log_radon_scale, county_effect_scale, and bias). The x𝑖 are features (log_uranium, floor_of_house, and floor_by_county) and the 𝑦𝑖 are target values (log_radon) for 𝑖 = 1…n observed data points:
VI is then characterized by:
(Technically we’re assuming q is absolutely continuous with respect to r. See also, Jensen’s inequality.)
Since the bound holds for all q, it is obviously tightest for:
Regarding terminology, we call
- q* the “surrogate posterior,” and,
- Q the “surrogate family.”
ω* represents the maximum-likelihood values of the deterministic parameters on the VI loss. See this survey for more information on variational inference.
Expressive surrogate posteriors
Next we estimate the posterior distributions of the parameters using VI with two different types of surrogate posteriors:
- A constrained multivariate Normal distribution, with covariance structure induced by a blockwise matrix transformation.
- A multivariate standard Normal distribution transformed by an Inverse Autoregressive Flow, which is then split and restructured to match the support of the posterior.
Multivariate Normal surrogate posterior
To build this surrogate posterior, a trainable linear operator is used to induce correlation among the components of the posterior.
We begin by constructing a base distribution with vector-valued standard Normal components, with sizes equal to the sizes of the corresponding prior components. The components are vector-valued so they can be transformed by the linear operator.
flat_event_size = tf.nest.map_structure(
tf.reduce_prod,
tf.nest.flatten(target_model.event_shape_tensor()))
base_standard_dist = tfd.JointDistributionSequential(
[tfd.Sample(tfd.Normal(loc=0., scale=1.), s)
for s in flat_event_size])To this distribution, we apply a trainable blockwise lower-triangular linear operator to induce correlation in the posterior. Within the linear operator, a trainable full-matrix block represents full covariance between two components of the posterior, while a block of zeros (or None) expresses independence. Blocks on the diagonal are either lower-triangular or diagonal matrices, so that the entire block structure represents a lower-triangular matrix.
Applying this bijector to the base distribution results in a multivariate Normal distribution with mean 0 and (Cholesky-factored) covariance equal to the lower-triangular block matrix.
operators = (
(tf.linalg.LinearOperatorDiag,), # Variance of uranium weight (scalar).
(tf.linalg.LinearOperatorFullMatrix, # Covariance between uranium and floor-by-county weights.
tf.linalg.LinearOperatorDiag), # Variance of floor-by-county weight (scalar).
(None, # Independence between uranium weight and county effects.
None, # Independence between floor-by-county and county effects.
tf.linalg.LinearOperatorDiag) # Independence among the 85 county effects.
)
block_tril_linop = (
tfp.experimental.vi.util.build_trainable_linear_operator_block(
operators, flat_event_size))
scale_bijector = tfb.ScaleMatvecLinearOperatorBlock(block_tril_linop)Finally, we allow the mean to take nonzero values by applying trainable Shift bijectors.
loc_bijector = tfb.JointMap(
tf.nest.map_structure(
lambda s: tfb.Shift(
tf.Variable(tf.random.uniform(
(s,), minval=-2., maxval=2., dtype=tf.float32))),
flat_event_size))The resulting multivariate Normal distribution, obtained by transforming the standard Normal distribution with the scale and location bijectors, must be reshaped and restructured to match the prior, and finally constrained to the support of the prior.
reshape_bijector = tfb.JointMap(
tf.nest.map_structure(tfb.Reshape, flat_event_shape))
unflatten_bijector = tfb.Restructure(
tf.nest.pack_sequence_as(
event_shape, range(len(flat_event_shape))))
event_space_bijector = target_model.experimental_default_event_space_bijector()Now, put it all together — chain the trainable bijectors together and apply them to the base standard Normal distribution to construct the surrogate posterior.
surrogate_posterior = tfd.TransformedDistribution(
base_standard_dist,
bijector = tfb.Chain([ # Chained bijectors are applied in reverse order.
event_space_bijector, # Constrain to the support of the prior.
unflatten_bijector, # Pack components into the event_shape structure.
reshape_bijector, # Reshape the vector-valued components.
loc_bijector, # Allow for nonzero mean.
scale_bijector # Apply the block matrix transformation.
]))Train the multivariate Normal surrogate posterior.
optimizer = tf.optimizers.Adam(learning_rate=1e-2)
@tf.function(jit_compile=True)
def run_vi():
return tfp.vi.fit_surrogate_posterior(
target_model.unnormalized_log_prob,
surrogate_posterior,
optimizer=optimizer,
num_steps=10**4,
sample_size=16)
mvn_loss = run_vi()
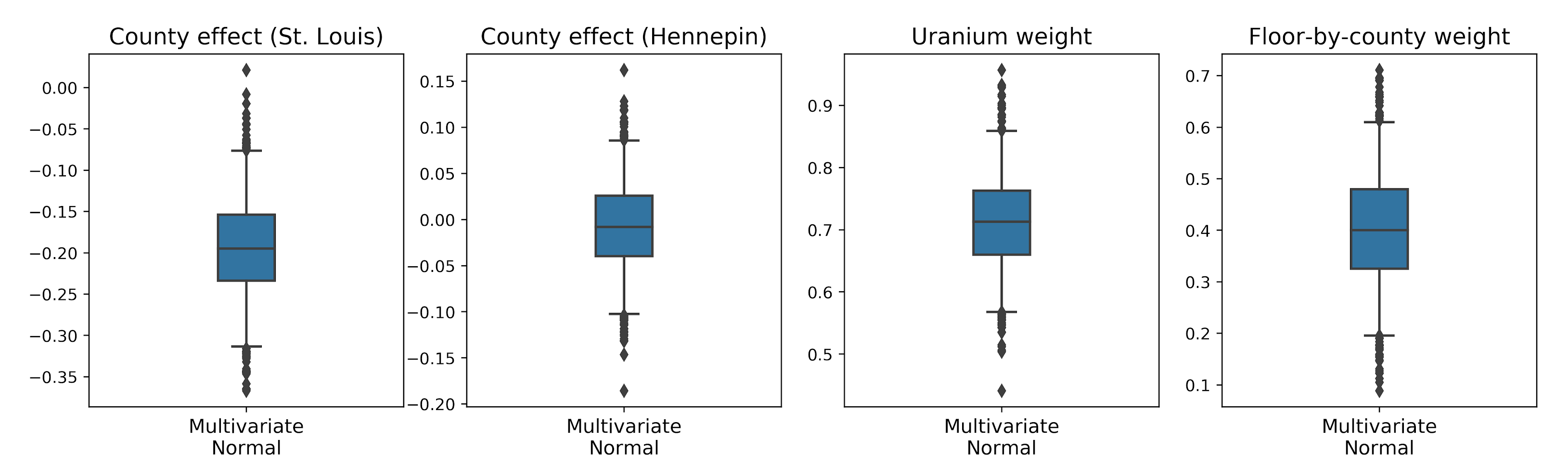
mvn_samples = surrogate_posterior.sample(1000)Since the trained surrogate posterior is a TFP distribution, we can take samples from it and process them to produce posterior credible intervals for the parameters.
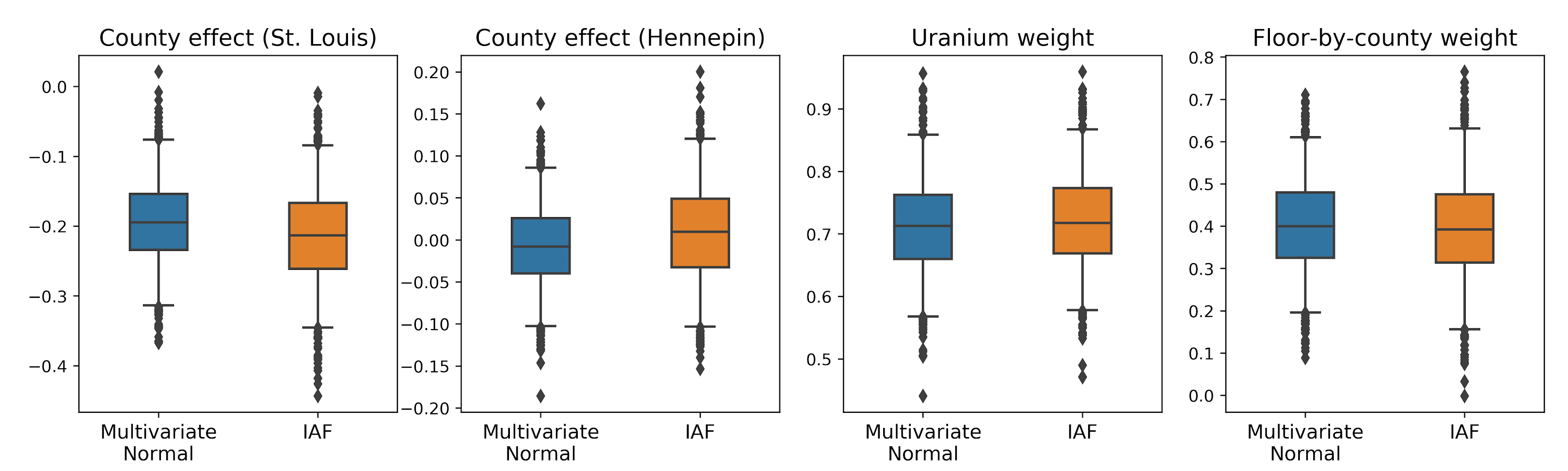
The box-and-whiskers plots below show 50% and 95% credible intervals for the county effect of the two largest counties and the regression weights on soil uranium measurements and mean floor by county. The posterior credible intervals for county effects indicate that location in St. Louis county is associated with lower radon levels, after accounting for other variables, and that the effect of location in Hennepin county is near neutral.
Posterior credible intervals on the regression weights show that higher levels of soil uranium are associated with higher radon levels, and counties where measurements were taken on higher floors (likely because the house didn’t have a basement) tend to have higher levels of radon, which could relate to soil properties and their effect on the type of structures built.
The (deterministic) coefficient of floor is -0.7, indicating that lower floors have higher radon levels, as expected.
Inverse autoregressive flow surrogate posterior
Inverse autoregressive flows (IAFs) are normalizing flows that use neural networks to capture complex, nonlinear dependencies among components of the distribution. Next we build an IAF surrogate posterior to see whether this higher-capacity, more flexible model outperforms the constrained multivariate Normal.
We begin by building a standard Normal distribution with vector event shape, of length equal to the total number of degrees of freedom in the posterior.
base_distribution = tfd.Sample(
tfd.Normal(loc=0., scale=1.),
sample_shape=[tf.reduce_sum(flat_event_size)])A trainable IAF transforms the Normal distribution.
num_iafs = 2
iaf_bijectors = [
tfb.Invert(tfb.MaskedAutoregressiveFlow(
shift_and_log_scale_fn=tfb.AutoregressiveNetwork(
params=2,
hidden_units=[256, 256],
activation='relu')))
for _ in range(num_iafs)
]The IAF bijectors are chained together with other bijectors to build a surrogate posterior with the same event shape and support as the prior.
iaf_surrogate_posterior = tfd.TransformedDistribution(
base_distribution,
bijector=tfb.Chain([
event_space_bijector, # Constrain to the support of the prior.
unflatten_bijector, # Pack components into the event_shape structure.
reshape_bijector, # Reshape the vector-valued components.
tfb.Split(flat_event_size), # Split into parts, same size as prior.
] + iaf_bijectors)) # Apply a flow model.Like the multivariate Normal surrogate posterior, the IAF surrogate posterior is trained using tfp.vi.fit_surrogate_posterior. The credible intervals for the IAF surrogate posterior appear similar to those of the constrained multivariate Normal.
Mean-field surrogate posterior
VI surrogate posteriors are often assumed to be mean-field (independent) Normal distributions, with trainable means and variances, that are constrained to the support of the prior with a bijective transformation. We define a mean-field surrogate posterior in addition to the two more expressive surrogate posteriors, using the same general formula as the multivariate Normal surrogate posterior. Instead of a blockwise lower triangular linear operator, we use a blockwise diagonal linear operator, in which each block is diagonal:
operators = (
tf.linalg.LinearOperatorDiag,
tf.linalg.LinearOperatorDiag,
tf.linalg.LinearOperatorDiag,
)
block_diag_linop = (
tfp.experimental.vi.util.build_trainable_linear_operator_block(
operators, flat_event_size))In this case, the mean field surrogate posterior gives similar results to the more expressive surrogate posteriors, indicating that this simpler model may be adequate for the inference task. As a “ground truth”, we also take samples with Hamiltonian Monte Carlo (see the Colab for the full example). All three surrogate posteriors produced credible intervals that are visually similar to the HMC samples, though sometimes under-dispersed due to the effect of the ELBO loss, as is common in VI.
Conclusion
In this post, we built VI surrogate posteriors using joint distributions and multipart bijectors, and fit them to estimate credible intervals for weights in a regression model on the radon dataset. For this simple model, more expressive surrogate posteriors appeared to perform similarly to a mean-field surrogate posterior. The tools we demonstrated, however, can be used to build a wide range of flexible surrogate posteriors suitable for more complex models.
Check out our code, documentation, and further examples on the TFP home page.