Ahmed Elnaggar and Michael Heinzinger are helping computers read proteins as easily as you read this sentence.
The researchers are applying the latest AI models used to understand text to the field of bioinformatics. Their work could accelerate efforts to characterize living organisms like the coronavirus.
By the end of the year, they aim to launch a website where researchers can plug in a string of amino acids that describe a protein. Within seconds, it will provide some details of the protein’s 3D structure, a key to knowing how to treat it with a drug.
Today, researchers typically search databases to get this kind of information. But the databases are growing rapidly as more proteins are sequenced, so a search can take up to 100 times longer than the approach using AI, depending on the size of a protein’s amino acid string.
In cases where a particular protein hasn’t been seen before, a database search won’t provide any useful results — but AI can.
“Twelve of the 14 proteins associated with COVID-19 are similar to well validated proteins, but for the remaining two we have very little data — for such cases, our approach could help a lot,” said Heinzinger, a Ph.D. candidate in computational biology and bioinformatics.
While time consuming, methods based on the database searches have been 7-8 percent more accurate than previous AI methods. But using the latest models and datasets, Elnaggar and Heinzinger cut the accuracy gap in half, paving the way for a shift to using AI.
AI Models, GPUs Drive Biology Insights
“The speed at which these AI algorithms are improving makes me optimistic we can close this accuracy gap, and no field has such fast growth in datasets as computational biology, so combining these two things I think we will reach a new state of the art soon,” said Heinzinger.
“This work couldn’t have been done two years ago,” said Elnaggar, an AI specialist with a Ph.D. in transfer learning. “Without the combination of today’s bioinformatics data, new AI algorithms and the computing power from NVIDIA GPUs, it couldn’t be done,” he said.
Elnaggar and Heinzinger are team members in the Rostlab at the Technical University of Munich, which helped pioneer this field at the intersection of AI and biology. Burkhard Rost, who heads the lab, wrote a seminal paper in 1993 that set the direction.
The Semantics of Reading a Protein
The underlying concept is straightforward. Proteins, the building blocks of life, are made up of strings of amino acids that need to be interpreted sequentially, just like words in a sentence.
So, researchers like Rost started applied emerging work in natural-language processing to understand proteins. But in the 1990s they had very little data on proteins and the AI models were still fairly crude.
Fast forward to today and a lot has changed.
Sequencing has become relatively fast and cheap, generating massive datasets. And thanks to modern GPUs, advanced AI models such as BERT can interpret language in some cases better than humans.
AI Models Grow 6x in Sophistication
The breakthroughs in natural-language processing have been particularly breathtaking. Just 18 months ago, Elnaggar and Heinzinger reported on work using a version of recurrent neural network models with 90 million parameters; this month their work leveraged Transformer models with 567 million parameters.
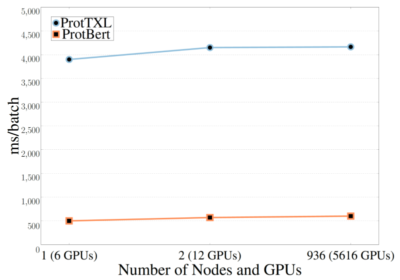
“Transformer models are hungry for compute power, so to do this work we used 5,616 GPUs on the Summit supercomputer and even then it took up to two days to train some of the models,” said Elnaggar.
Running the models on thousands of Summit’s nodes presented challenges.
Elnaggar tells a story familiar to those who work on supercomputers. He needed lots of patience to sync and manage files, storage, comms and their overheads at such a scale. He started small, working on a few nodes, and moved a step at a time.
“The good news is we can now use our trained models to handle inference work in the lab using a single GPU,” he said.
Now Available: Pretrained AI Models
Their latest paper, published in July, characterizes the pros and cons of a handful of the latest AI models they used on various tasks. The work is funded with a grant from the COVID-19 High Performance Computing Consortium.
The duo also published the first versions of their pretrained models. “Given the pandemic, it’s better to have an early release,” rather than wait until the still ongoing project is completed, Elnaggar said.
“The proposed approach has the potential to revolutionize the way we analyze protein sequences,” said Heinzinger.
The work may not in itself bring the coronavirus down, but it is likely to establish a new and more efficient research platform to attack future viruses.
Collaborating Across Two Disciplines
The project highlights two of the soft lessons of science: Keep a keen eye on the horizon and share what’s working.
“Our progress mainly comes from advances in natural-language processing that we apply to our domain — why not take a good idea and apply it to something useful,” said Heinzinger, the computational biologist.
Elnaggar, the AI specialist, agreed. “We could only succeed because of this collaboration across different fields,” he said.
See more stories online of researchers advancing science to fight COVID-19.
The image at top shows language models trained without labelled samples picking up the signal of a protein sequence that is required for DNA binding.
The post Learning Life’s ABCs: AI Models Read Proteins to Fight COVID-19 appeared first on The Official NVIDIA Blog.
